|
MVLib - Short Writeup - Version 5.0 Statistic, Signal filters, Fourier components - Last update: March 20, 2014 Source code available for local users only |
|||||||||||||||||||||||||||||||||||
| In the second column of the following table, S stays for "Subroutine", I stays for "Integer function" and R stays for "Real function". | |||||||||||||||||||||||||||||||||||
| average(x,n) | R | Returns the average of the first n components of vector v. | |||||||||||||||||||||||||||||||||
| sigma2(x,n) | R | Returns the variance of the first n components of vector v. | |||||||||||||||||||||||||||||||||
| bias(x,y,n) | R |
Returns the bias of the first n components of vectors
x and y 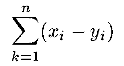 . .
|
|||||||||||||||||||||||||||||||||
| rmst(x,y,n) | R |
Returns the Root Mean Square
of the first n components of vectors x and y
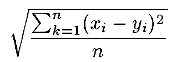 . .
| |||||||||||||||||||||||||||||||||
| rmstn(x,y,n) | R |
Returns the Normalized Root Mean Square
of the first n components of vectors x and y
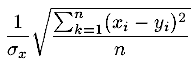 . .
|
|||||||||||||||||||||||||||||||||
| vcorr(x,y,n) | R |
Returns the absolute value of linear correlation coefficient
of the first n components of vectors x and y,
the coeffient ranges from 1 for a perfect correlation (or
anticorrelation) to 0
for no correlation and is calculated as
the absolute value of the following expression:
 . .
|
|||||||||||||||||||||||||||||||||
| wcorr(x,y,n) | R |
Returns the linear correlation coefficient
of the first n components of vectors x and y,
the coeffient ranges from 1 for a perfect correlation to -1
for a perfect anticorrelation and is calculated as:
 . .
|
|||||||||||||||||||||||||||||||||
| rainscore(x,y,n,s,out) | S |
Returns different statistic parameters for the first n
components of two precipitation time series x and y,
the real input argument s specifies the threshold and the
results are returned in the components of vector out.
|
|||||||||||||||||||||||||||||||||
| nashsutcliffe(qo,qm,n) | R |
Returns the Nash-Sutcliffe model efficiency coefficient. This
coefficient is used to assess the predictive power of hydrological models.
It is defined as: 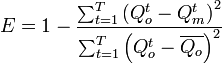 See also: Nash, J. E. and J. V. Sutcliffe (1970), River flow forecasting through conceptual models part I - A discussion of principles, Journal of Hydrology, 10 (3), 282-290. |
|||||||||||||||||||||||||||||||||
| cioppskill(qo,qm,n) | R | Old version of nashsutcliffe routine documented above. This entry point is saved for backward compatibility only. | |||||||||||||||||||||||||||||||||
| ttest(d1,n1,d2,n2) | R |
Returns the T-test (or student test) to check the significancy of
differences between two distributions, calculated as: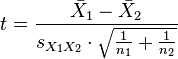 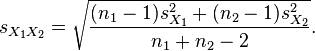 |
|||||||||||||||||||||||||||||||||
| ttestthresh(n,sigma) | R | Returns the threshold value to be considered to evaluate the T-test (see routine ttest above), the first argument is the number of degree of freedom of distribution, the second real argument is the significancy level and the allowed values are: 0.9995, 0.999, 0.9975, 0.995, 0.99, 0.975, 0.95 and 0.90. Data are taken from the table reported at Wikipedia page. | |||||||||||||||||||||||||||||||||
| Functions | |||||||||||||||||||||||||||||||||||
| sigmoide(x,a,b) | R | Returns the value of the sigmoid in the abscissa specified by the first argument; the second argument specifies the center of the sigmoid, the third argument specifies the distance from the center at which the sigmoid value is 0.1/0.9, namely the sigmoid is 0.1 when x=a-b and the sigmoid is 0.9 when x=a+b. | |||||||||||||||||||||||||||||||||
| Random extraction | |||||||||||||||||||||||||||||||||||
| acaso(xx) | R | Returns a random number with a flat distribution in the range (0-1). The algorithm is platform independent and can be initialized using the MVLib parameter Random Seed. xx is a dummy argument. | |||||||||||||||||||||||||||||||||
| gauss(average,sigma) | R | Returns a random number with a gaussian distribution. average is the average value and sigma is the standard deviation. The algorithm is platform independent and can be initialized using the MVLib parameter Random Seed. | |||||||||||||||||||||||||||||||||
| Fourier Tranform | |||||||||||||||||||||||||||||||||||
| mvft(d,nd,a,b,n) | S |
Calculates the first n coefficients of Fourier series for
the first nd components of cevctor d, following:
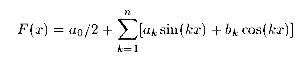 . The results
are returned in the vectors a(0:n-1) and b(0:n-1).
By definition is b(0)=0. . The results
are returned in the vectors a(0:n-1) and b(0:n-1).
By definition is b(0)=0.
|
|||||||||||||||||||||||||||||||||
| mvftinv(d,nd,a,b,n) | S | mvftinv carryout the inverse Fourier tranform. The first nd components of cevctor d are rebuilt using the n coefficients of Fourier series stored in the vectors a(0:n-1) and b(0:n-1). | |||||||||||||||||||||||||||||||||
| mvftfilt(in,ou,n,ispett1,ispett2) | S | mvftfilt is medium frequencies filter. It rebuild in the vector out the n components of vector in, considering only the Fourier component between ispett1 and ispett2. | |||||||||||||||||||||||||||||||||
| mvfthffilter(inp,out,n,a,b,c,w) | S |
Allows to selecty the high frequencies in the n components
of input real vector inp, the results are returned in the
real vector out. The real vectors a and b will
contain, as output, the smoothed Fourier coefficients. These
coefficients are smoothed following a sigmoid function whose
characteristic are specified by the real arguments c
and w: the first specify the center of the sigmoid, the latter
specify the width of the function, namely the value for which the
sigmoid si smoothed by a factor 0.1/0.9; as an example if
the arguments are n=200, c=100.0 and w=30.0,
the component number 100 (the center of the sigmoide range)
will be smoothed by a factor 0.5, the component number 130
will be smoothed by a factor 0.9 and the component number 70
will be smoothed by a factor 0.1. The
sigmoid is calculated as follow:
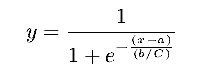
| |||||||||||||||||||||||||||||||||
| mvfthffilter(inp,out,n,a,b,c,w) | S | Same as mvfthffilter but in this case low frequencies are selected, the sigmoid is this. | |||||||||||||||||||||||||||||||||
| Signal filters | |||||||||||||||||||||||||||||||||||
| naverage (x,y,m,n) | S |
The input vector x of m components is filtered with a
n-average approach. The output vector y is calculated as
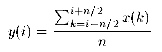
| |||||||||||||||||||||||||||||||||
| o1derivative(x,y,n) | S |
The input vector x of n components is filtered with a
first order derivative. The output vector y is calculated as
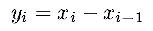
medianfilter(x,y,n,m) |
S |
The input vector x of n components is filtered with a
median value calulated over m components. The result is returned in the real vector y.
| dewmidnightfilter(y,ndata,noval,title) |
S |
Allows to filter the so called midnight bug of dewetra
precipitation data. In many circumstances, the midnight data corresponds to the accumulated rain in the last N hours and N is often (but not always) 24. When the
midnight value is anomalous respect the to values in the last 23 hours, the midnight value is set to no-data. | The first argument is the time series, the second integer argument is the number of data (usually 8760 or 8784), the third integer argument is the no-data value (usually -9999), the last argument is a string, if this argument is not empty a log is produced by this routine.
d1cellcycle(vec,rule,dummy,n) |
S |
See the Cellular Automata routines description
|
cavectorfill(v,w,z,ndat) |
S |
See the Cellular Automata routines description
|
| 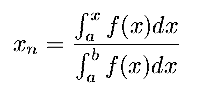 This is usually referred to as quantic normalization.
This is usually referred to as quantic normalization.
qnormdef(id,v,n) |
S |
The input vector x of n components is used to define the
parameters of quantic normalization. The integer input flag id
is not yet used, it can be used in the next versions to manipulate
several normalization within the same application.
|
qnormal(id,x) |
R |
Returns the normalized value of input real argument x.
The integer input flag id is not yet used.
|
qanormal(id,x) |
R |
Returns the annormalized value of input real argument x.
The integer input flag id is not yet used.
|
Time series analisys
| autocorrelation(x,n,y,lags) |
S |
Calculates the autocorrelation between the components of real vector x(n)
considering an interval of 1-lags elements. The first component of
output vector y will return the linear correlation between the elements
x(i) and elements x(i+1); the k-th component of
output vector y will return the linear correlation between the elements
x(i) and elements x(i+k).
|
autocorrplot(x,n,y,lags) |
S |
Same as autocorrelation routine, but an histogram is also produced.
|
| ||||